This week, I had a quick sync with the mentors as it seemed like I couldn't derive systematic results from running my previous experiments.
I liked the idea to introduce some noise to some dataset that has a low rate (e.g. less than 1-3% of misclassified labels to compare baseline with cleanlab. I used Fashion MNIST and introduced some random corruption to the training set by flipping the labels.
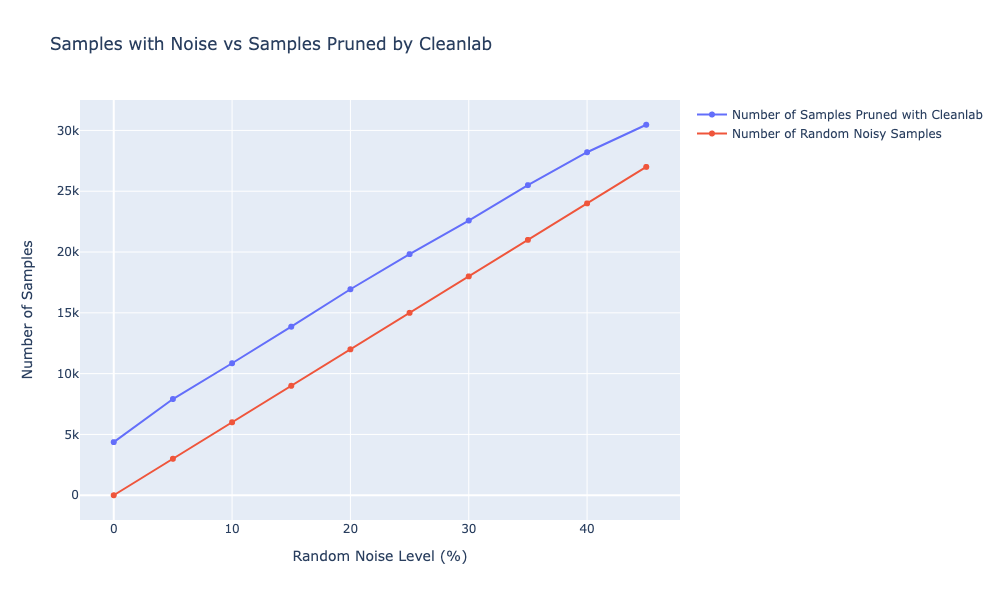
In one of my experiments, I set the maximum noise to 50% and gradually introduced 5% the noise at each step, comparing the performance of baseline and cleanlab in parallel. This time, instead of relabelling, I decided to prune the samples with low confidence scores. Here’s a quick example: if I have 60,000 samples in the dataset, at 10% of the noise, I’d randomly flip 6,000 labels. The baseline would then be trained with all samples (60,000), but the cleanlab would be trained only on the labels that weren’t classified as erroneous by cleanlab . For example, If cleanlab found that 5,000 are labeled incorrectly, then I would only use 55,000 images for training.
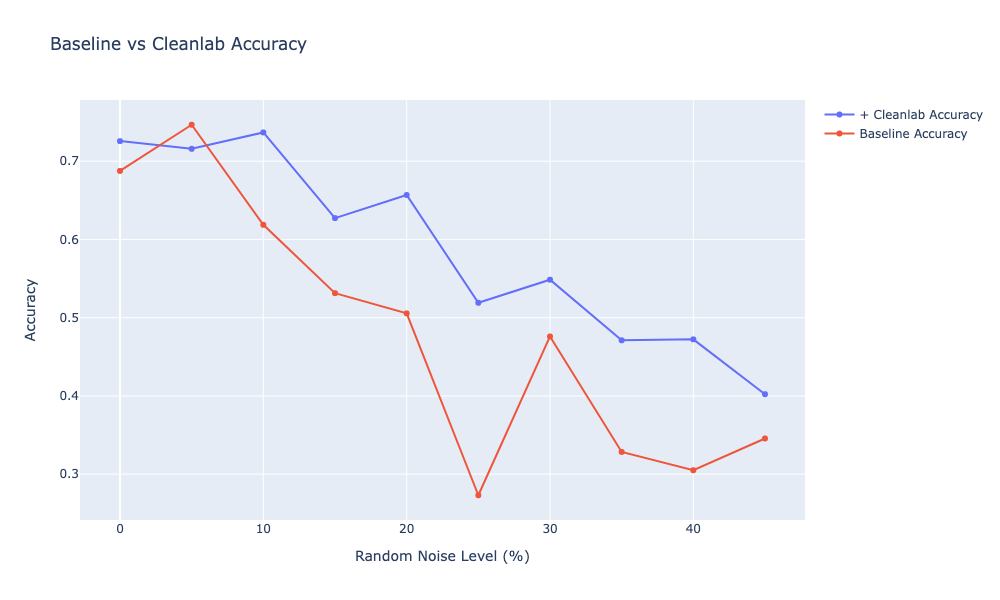
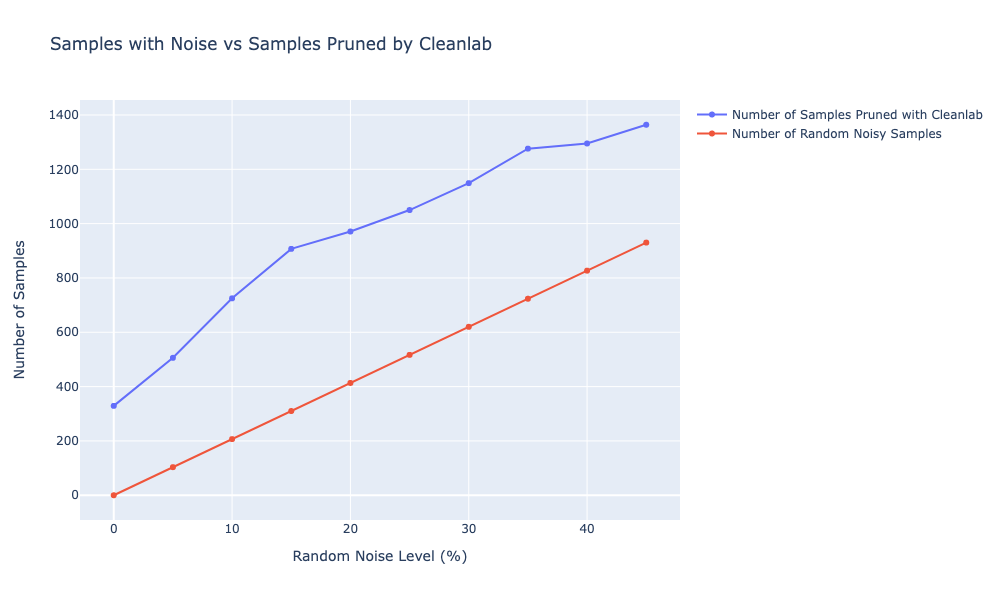
It seems that pruning the samples with lower confidence works well, as cleanlab seems to remove the labels that were introduced with each noise level. We can also see that the accuracy stays around 80% with cleanlab, while with the random noise (e.g. without removing any samples) it drops linearly. On average, I can also see that cleanlab on average prunes more labels than I initially introduced in the data. The mean of additional samples that cleanlab discards is ≈4500 samples across all noise levels. Since I don’t know the true noise in the original dataset, it’s hard to say whether cleanlab is doing a good job on removing these, but I would argue that cleanlab seems to be overestimating. However, it seems to systematically pick up the newly introduced noisy labels and identify them as erroneous.
I was surprised that after introducing random noise, CL would still prune the erroneous labels. The way CL algorithm works is by accurately and directly characterizing the uncertainty of label noise in the data. The foundation CL depends on is that label noise is class-conditional, depending only on the latent true class, not the data . For instance, a leopard is likely to be mistakenly labeled as a jaguar . Cleanlab takes that assumption and computes joint distribution among different classes (e.g. 3% of the data is labeled leopard (noisy label), but the true label is jaguar). The main idea is that underlying data has implications for the labeler’s decisions, and I basically took this assumption out of equation after randomly swapping labels in the dataset (e.g. I didn’t care if a certain class would be more likely to be mislabelled as another class). I would think that CL algorithm relies on this assumption heavily to guess which label to prune, but the performance was still accurate and stable. In the real-world noisy data, the mislabelling between different classes would have a stronger statistical dependence, so I can say that this example was even a bit more difficult for cleanlab.
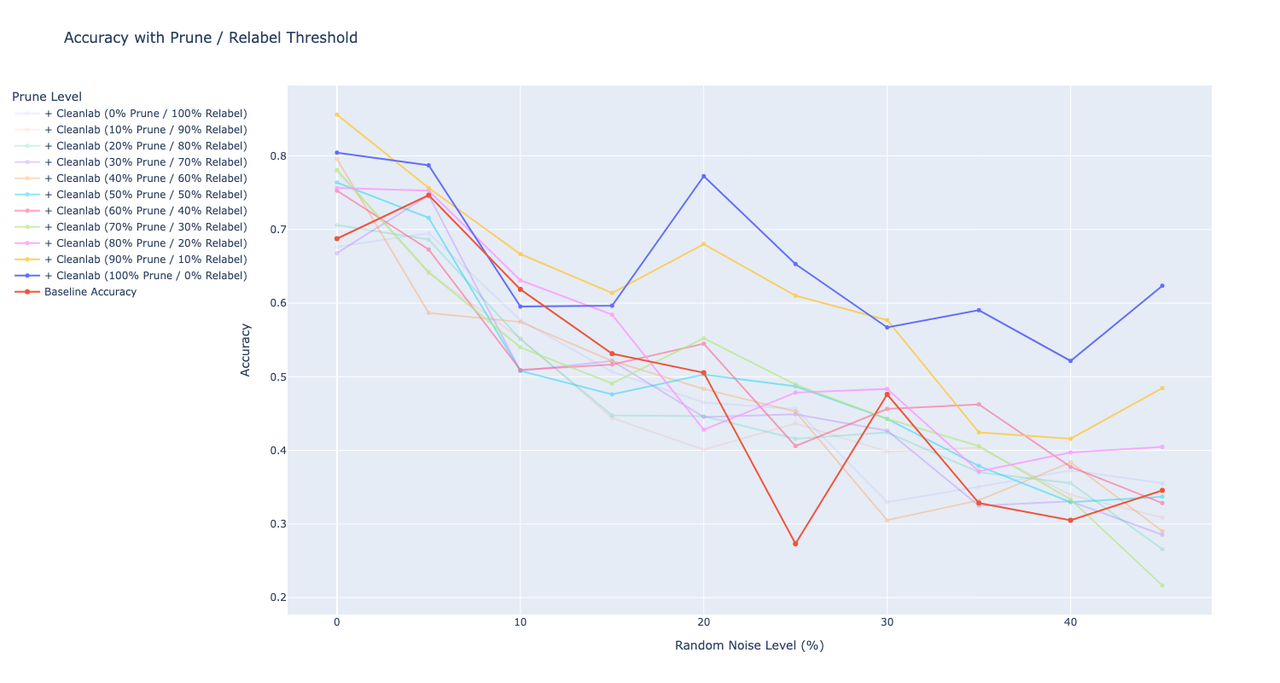
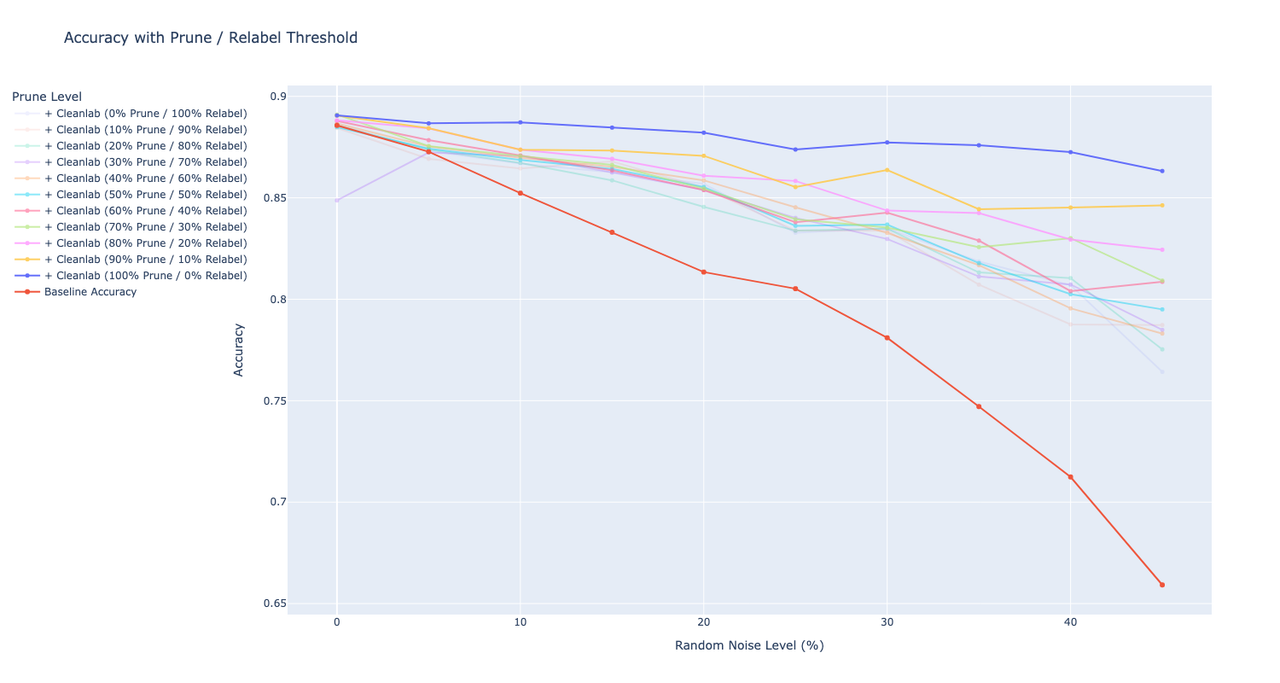
I’ve tried to experiment with different threshold values for pruning and relabelling the images (e.g. remove 20% of the images with lowest label quality, but leave and relabel the rest). I’ve started with a threshold of 0% (e.g. relabel all labels to the ones predicted by cleanlab ) and then gradually increased the threshold value with a 10% step till I reached 100% prune level (e.g. remove all labels that were found to be erroneous by cleanlab). As before, I run these from 0 to 50% of the noise level.On the graph, I plotted the accuracy of the models trained with training sets that were fixed with different threshold values. For example, 100% Prune / 0% Relabel indicates the accuracy of the model when all erroneous samples and their labels were deleted, while 0% Prune / 100% Relabel shows the accuracy of the model when all of the samples were left but relabelled.Looking at the graph, I can say that cleanlab definitely does a great job at identifying labels, but not necessarily at fixing them automatically. As soon as I increase the % of labels that I’d like to relabel, the accuracy starts to go down in linear way. The training set with 100% of pruning got the highest accuracy, while the training set with all labels relabelled got the worst accuracy on the fixed model.As a next step, I can try to see what happens if we only remove a certain % of erroneous samples, but leave the labels of the other erroneous samples as they are. (edited)
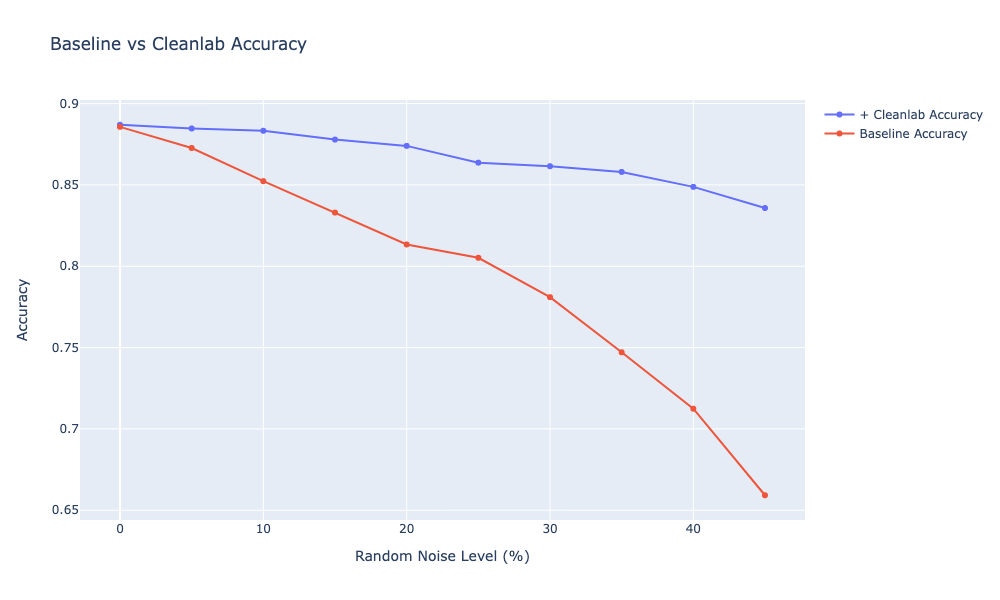
I also run this pipeline on the Roman Dataset (DCAI). This dataset is quite noisy, so there’s not a ton of improvement on 0-10% of the noise level. However, as I introduce more noisy labels, looks like cleanlab is still able to pick them up. Running a few more trials to see what’s the performance with different Prune vs Relabel threshold.